
April 2, 2001
New images reveal the workings of cellular protein factories
By Tim Stephens
The workings of a tiny molecular machine crucial to all forms of life are emerging
from highly detailed new images obtained by researchers at the University of California,
Santa Cruz. In a paper published by the journal Science on March 30, the researchers
describe the structure of the ribosome, a complex particle just one millionth of
an inch in diameter, in sufficient detail to begin to understand how it works.
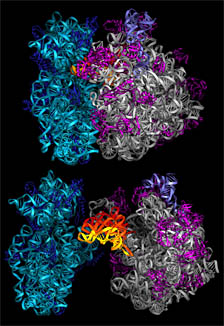 |
| New images are adding to scientists' understanding of ribosomes, the protein factories of all living cells. Image: Marat Yusupov et al. |
Ribosomes are the protein factories of all living cells. They hold the equipment necessary to read the genetic code and translate it into specific protein structures. Inside every cell, tens of thousands of ribosomes churn out proteins with mind-boggling speed and precision.
They are ancient structures that show little variation among different forms of
life. Most research has focused on bacterial ribosomes, which are composed of three
different RNA molecules and more than 50 different proteins arranged in two major
subunits.
One of the most striking features of ribosomes is that the components that carry
out protein synthesis are made of RNA, a type of molecule similar in structure to
DNA. In contrast, the enzymes that catalyze most of the chemical reactions necessary
for life are proteins. The central role of RNA in the function of the ribosome is
an idea long championed by Noller and others, but only recently confirmed by a series
of landmark studies by Noller's group at UC Santa Cruz and by other researchers,
including Peter Moore and Thomas Steitz of Yale University. This month, the three
researchers will share the prestigious Lewis S. Rosenstiel Award for Distinguished
Work in Basic Medical Science in recognition of this important discovery. The new
paper provides further confirmation that ribosomal RNA is the active component in
protein synthesis.
"Why these ancient organelles use RNA, instead of protein, for the complex and
biologically crucial task of protein synthesis is a fascinating question, the answer
to which may shed light on the origins of life on Earth," Noller said.
Noller and his coworkers published the first detailed images of a complete ribosome
in 1999. In their new paper, the three-dimensional structure of the ribosome is revealed
in much finer detail, enabling them to identify specific components that carry out
key functions. The lead authors of the study are Marat Yusupov and Gulnara Yusupova,
visiting researchers at UCSC's Center for Molecular Biology of RNA, now at the French
national research center in Strasbourg. The coauthors, in addition to Noller, are
Albion Baucom and Kate Lieberman at UCSC; Thomas Earnest at Lawrence Berkeley National
Laboratory; and Jamie Cate, now at the Whitehead Institute in Cambridge, MA.
Biologists have known the basic outlines of protein synthesis for decades. The instructions
for making a protein are carried to the ribosome by a messenger RNA molecule, which
has copied the instructions from chromosomal DNA, the storehouse of genetic information
carried in almost every cell. The building blocks of proteins are carried to the
ribosome by transfer RNA molecules. On the ribosome, the transfer RNAs recognize
specific sequences of genetic code on the messenger RNA and line up the protein building
blocks in the proper order. The ribosome then catalyzes the formation of chemical
bonds between the building blocks.
The images obtained by Noller's group show not only the ribosome itself but also
messenger RNA and transfer RNAs in the positions they occupy during the process of
protein synthesis. Most of the action involved in protein synthesis seems to take
place at the interface between the two ribosomal subunits. Noller and his coworkers
identified 12 chemical bridges linking the two subunits, mostly involving ribosomal
RNA. They also located the sites where the ribosome interacts with the transfer RNAs.
These findings enabled them to propose mechanisms by which protein synthesis is coupled
with movements of specific components of the ribosome.
"The ribosome is a molecular machine, and it must have moving parts to accomplish
its function. We are now in a position to understand the structural rearrangements
of the ribosome during protein synthesis," Noller said.
This achievement has practical significance because many antibiotics work by binding
to and disrupting bacterial ribosomes. Understanding the ribosome's structure may
lead to the development of new and more effective antibiotics.
The images were obtained using a technique called x-ray crystallography. The improved
resolution of these new images is the result of fine-tuning some two dozen variables
in the group's experiments, Noller said. The ribosome is the largest molecular structure
ever solved by x-ray crystallography.